Model diagnostics
Linear Regression Assumptions: L-I-N-E
- L: Linear relationship between (Y) and the explanatory variable (X)
- I: Independence of errors—there’s no connection between how far any two points lie from the regression line
- N: Normal distribution of Y at each level of X
- E: equality of variance of the errors – variability remains the same for all levels of X.
In other words, the residuals (errors) should satisfy the following:
- L: The mean value for Y at each level of X lies on regression line.
- I: There is no clear pattern in the errors
- N: At each level of X, the values for Y are normally distributed.
- E: The variability in the Y’s for each level of X is the same
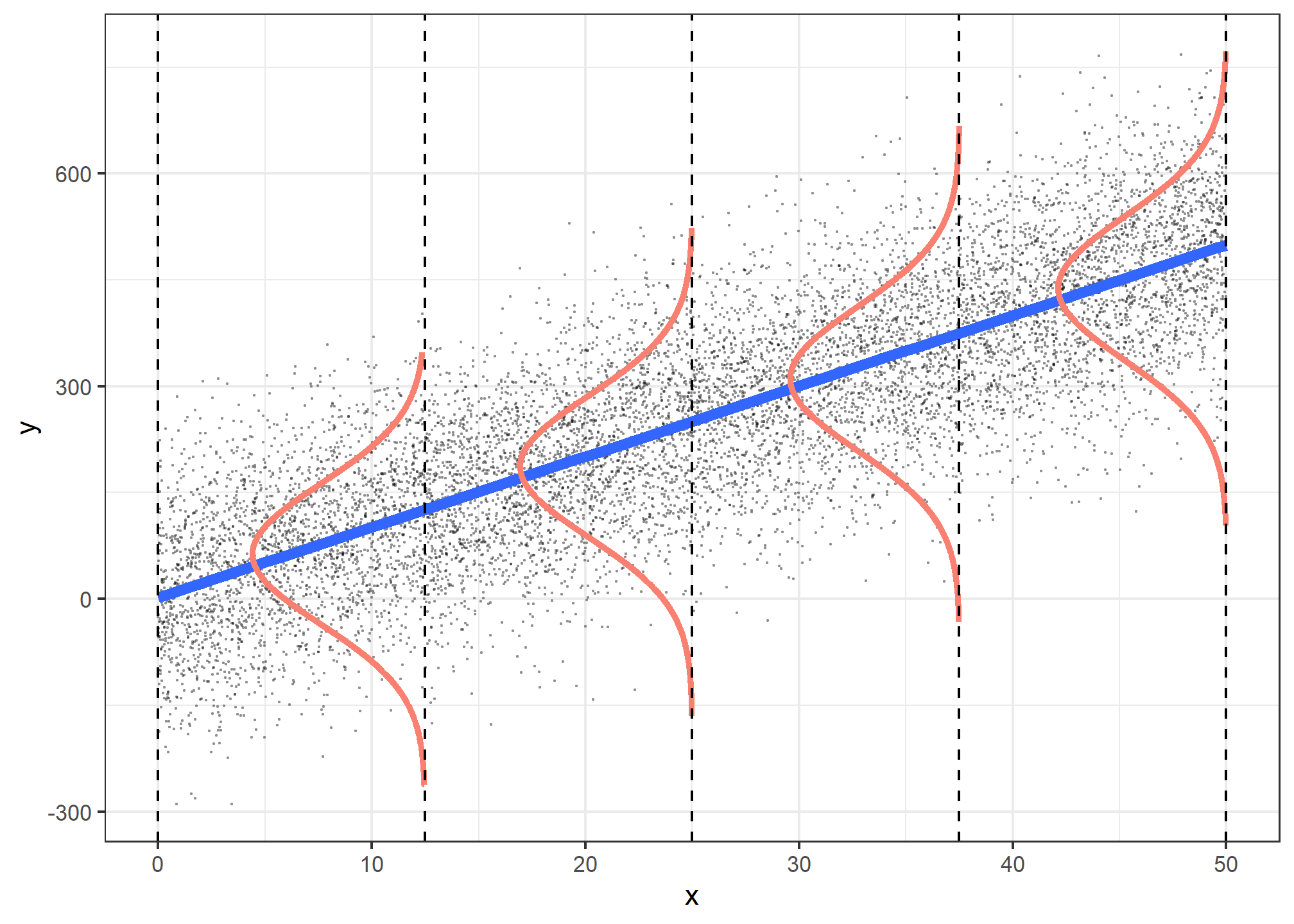
Figure 1: Assumptions for linear ordinary least squares (OLS) regression
Regression diagnostic plots with ggfortify::autoplot()
Let us see what is happening in our models. We will use the ggfortify package and its autoplot() command to get the following regression diagnostic plots:
- Residuals vs. Fitted: checks Linearity assumption. Residuals should be random, with no pattern, and around Y = 0; if not, there is a pattern in the data that is currently unaccounted for.
- Normal Q-Q: checks residual Normality assumption. Deviations from a straight line indicate that residuals do not follow a Normal distribution.
- Scale-Location: checks whether residuals have equal/constant variance or not. Positive or negative trends across the fitted values indicate variability that is not constant.
- Residuals vs. Leverage: check for influential points. Points with high leverage (having unusual values of the predictors) and/or high absolute residuals can have an undue influence on estimates of model parameters.
model1 <- lm(body_mass_g ~ flipper_length_mm, data = penguins)
model2 <- lm(body_mass_g ~ flipper_length_mm + species , data = penguins)
model3 <- lm(body_mass_g ~ flipper_length_mm + species + sex , data = penguins)
model4 <- lm(body_mass_g ~ flipper_length_mm + species + sex , data = penguins)
model5 <- lm(body_mass_g ~ flipper_length_mm + species + sex + bill_length_mm + bill_depth_mm , data = penguins)
model6 <- lm(body_mass_g ~ . , data = penguins)
library(ggfortify)
autoplot(model1) +
theme_minimal() +
labs (title = "Model 1 Diagnostic Plots")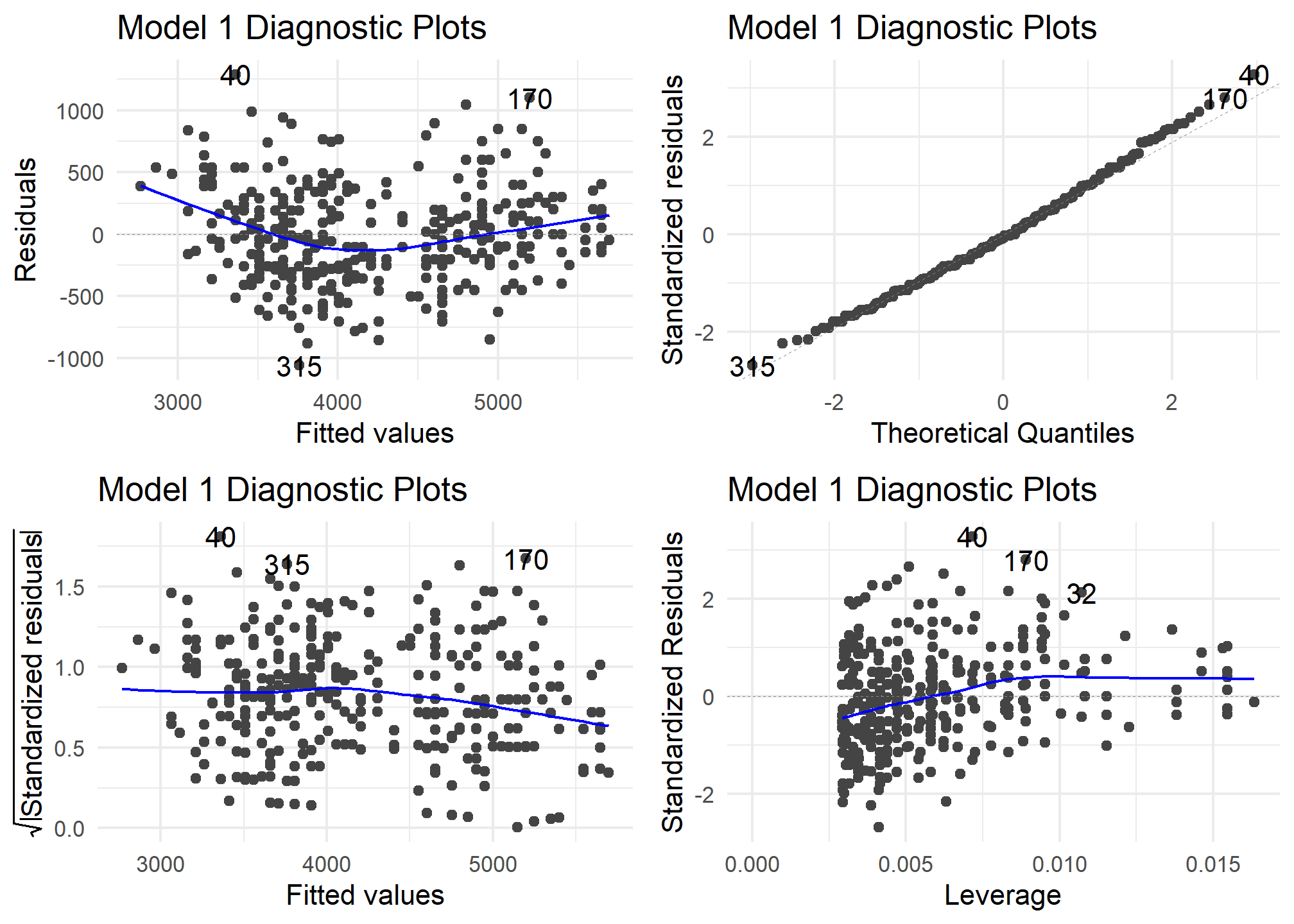
autoplot(model2) +
theme_minimal() +
labs (title = "Model 2 Diagnostic Plots")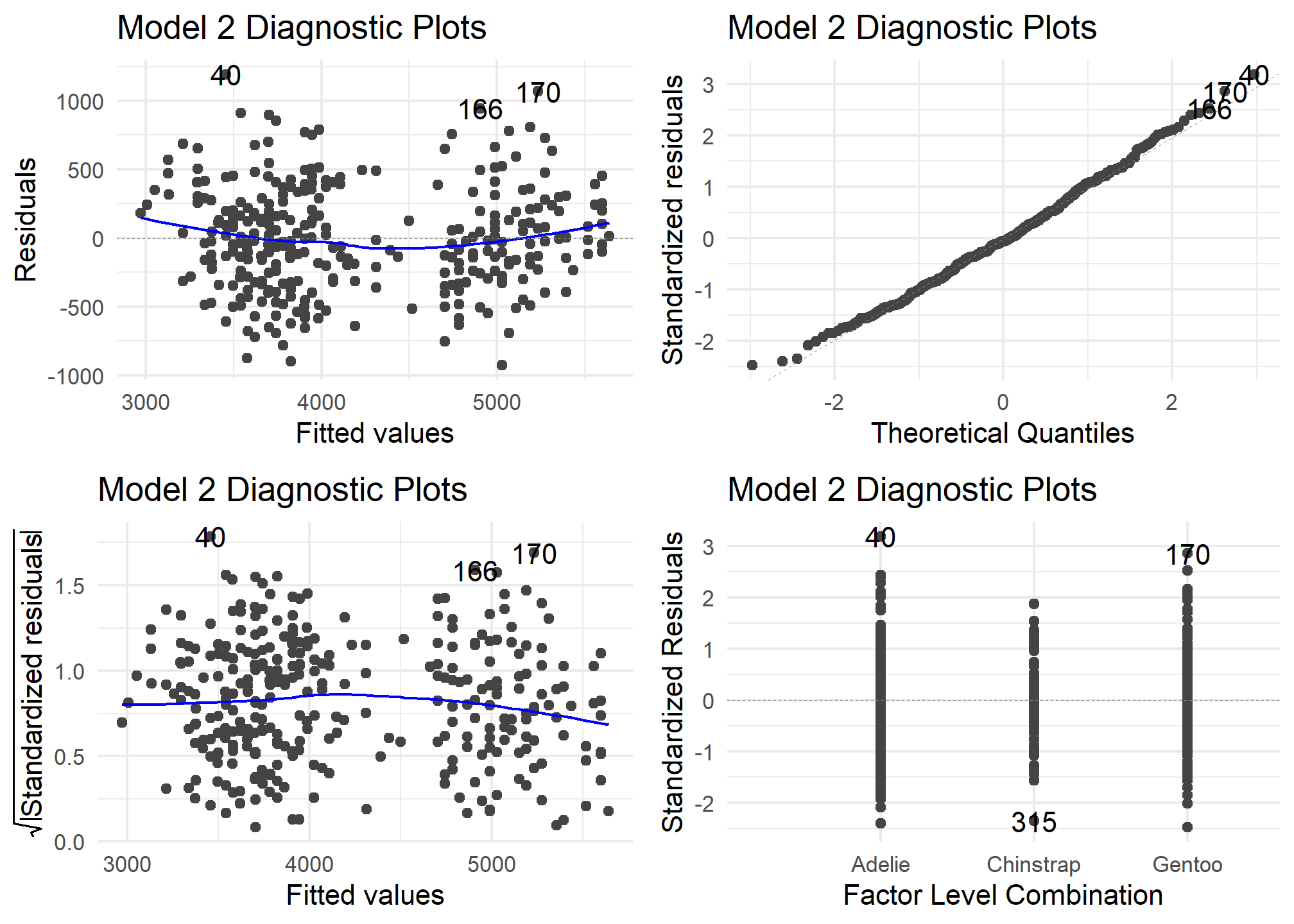
autoplot(model3) +
theme_minimal() +
labs (title = "Model 3 Diagnostic Plots")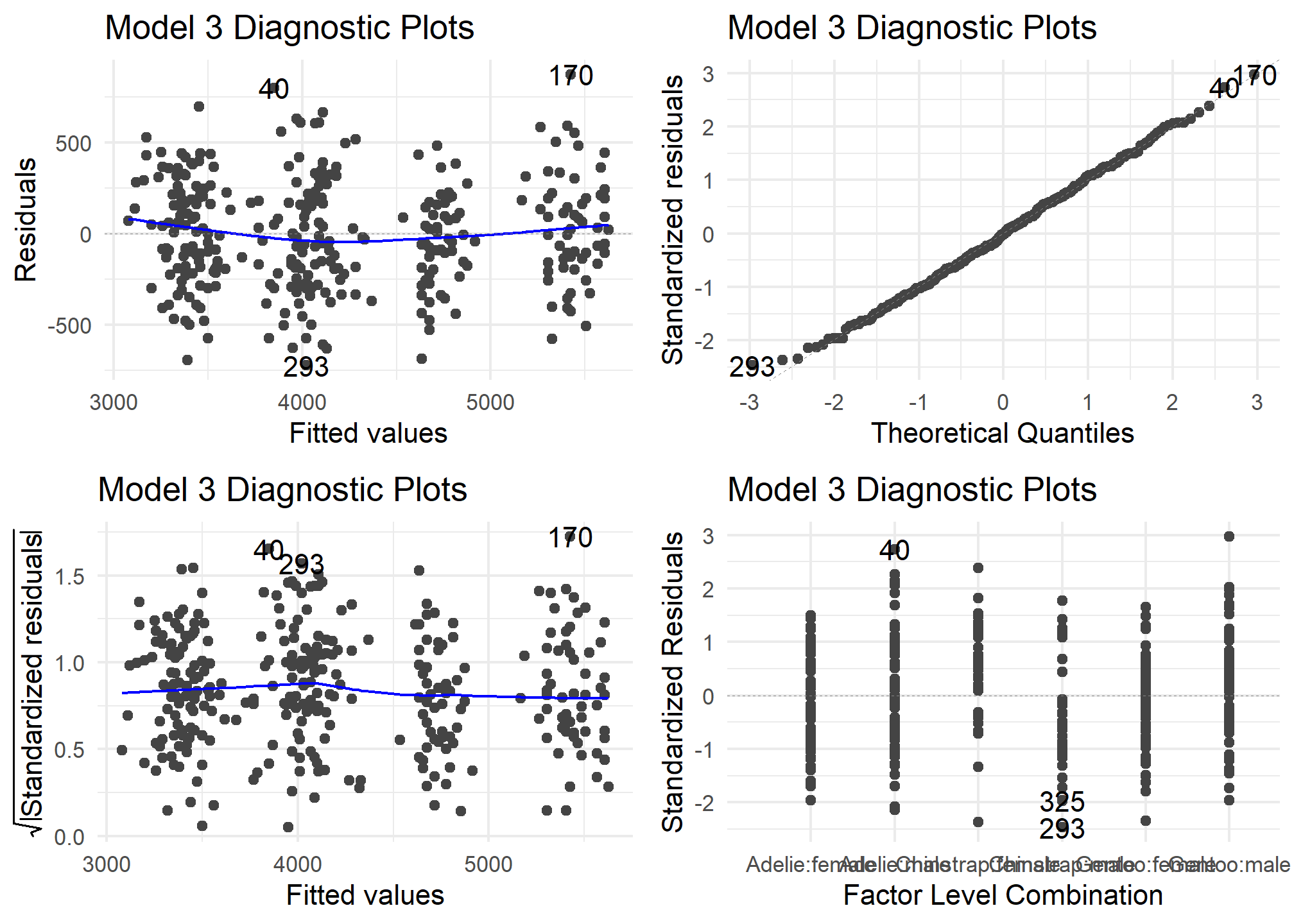
autoplot(model4) +
theme_minimal() +
labs (title = "Model 4 Diagnostic Plots")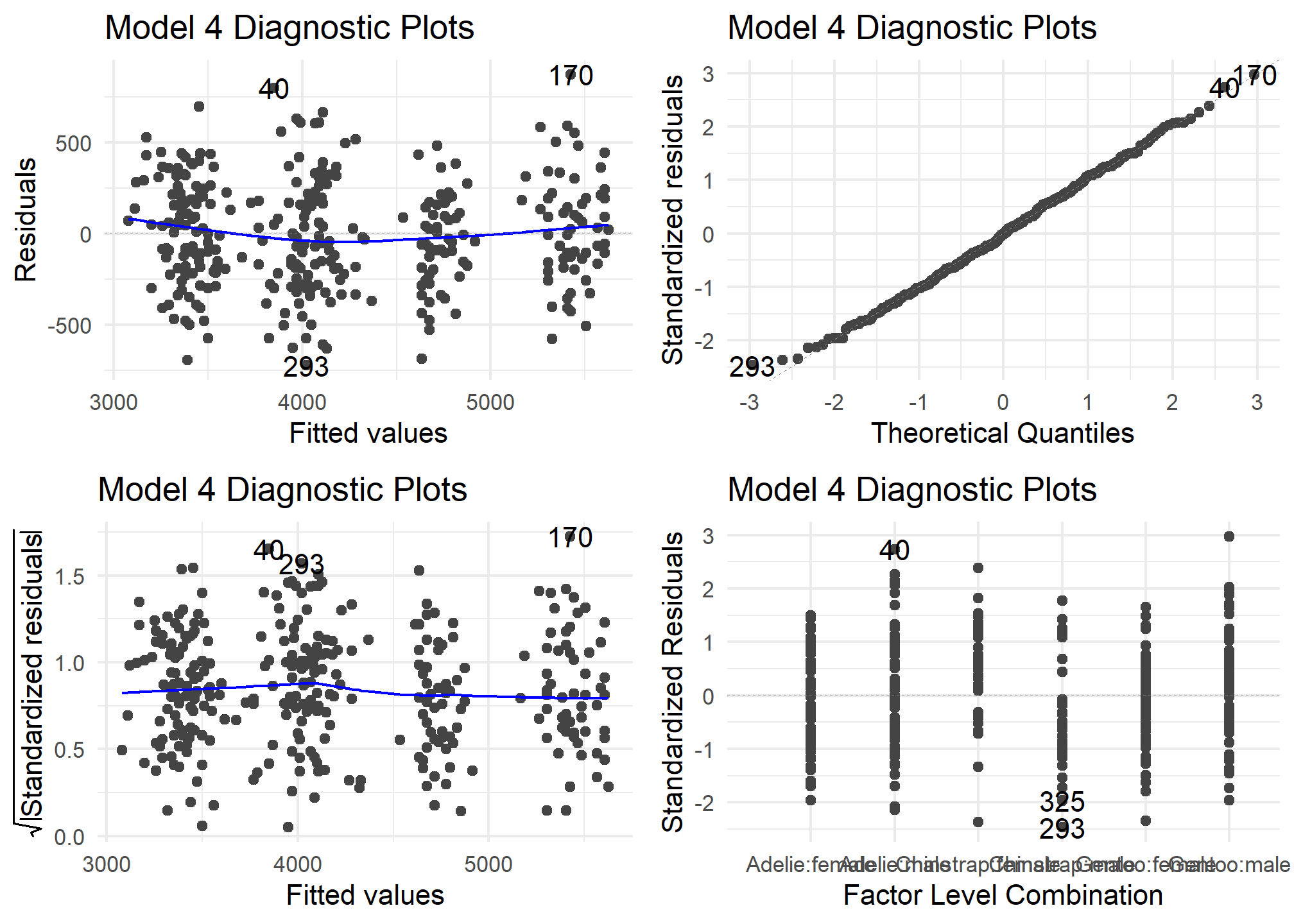
autoplot(model5) +
theme_minimal() +
labs (title = "Model 5 Diagnostic Plots")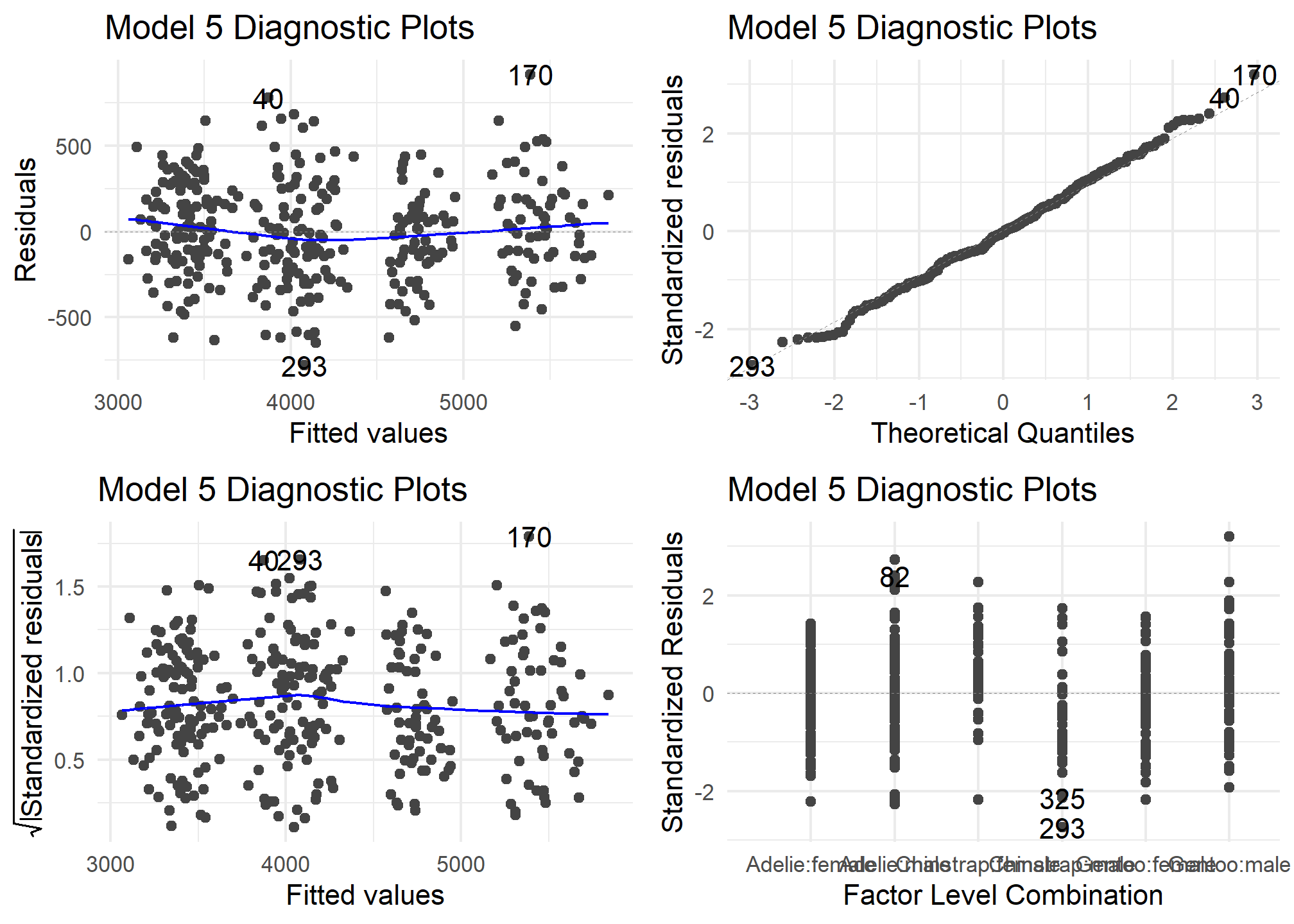
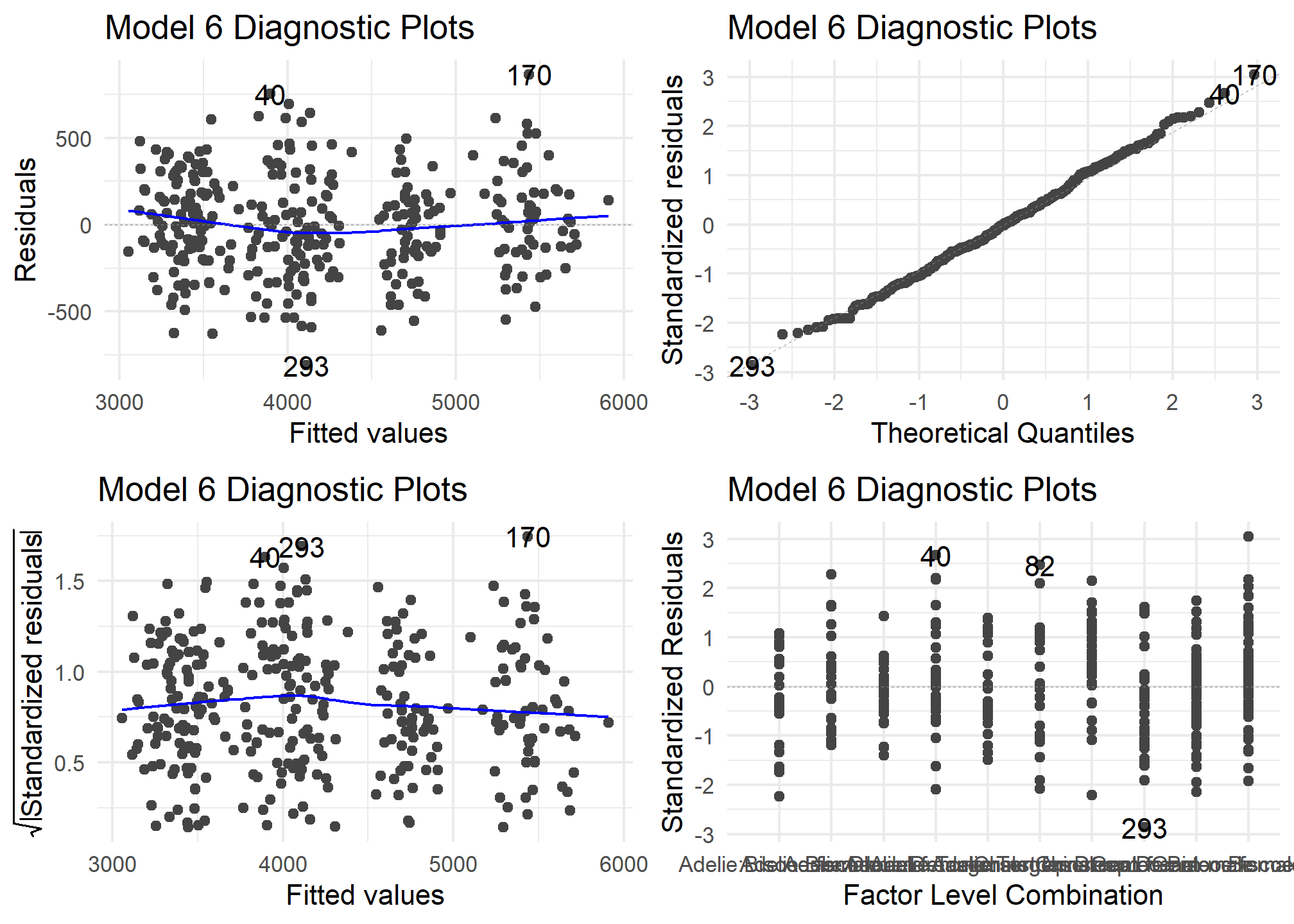
autoplot(model6) +
theme_minimal() +
labs (title = "Model 6 Diagnostic Plots")