Import data
Learning Objectives
1. Load external data from a .csv file into a data frame.
2. Describe what a data frame is.
3. Use indexing to subset specific portions of data frames.
4. Describe what a factor is.
5. Reorder and rename factors.
6. Format dates.
Overview
One of the things that I found strange when I started working with R was that, unlike other software like Excel, Stata, SPSS, etc., you couldn’t just double click on an .xls, .dta, or .sav file, load the data and look at its contents. In R, we must use a command to explicitly import the data into memory.
While there are many possible data formats, we will concentrate on
CSV files, namely Comma Separated Values files that are a common
way to save the raw data from spreadsheets, without any of the
formatting, etc. The readr R package contains functions for
importing data saved as flat file documents; readr is a core member
of the tidyverse and is loaded every time you call library(tidyverse).
CSV file names end with a .csv and if you opened one inside Excel, it would look like a regular Excel file. NASA provides an estimate of global surface temperature change which allows us to calculate weather anomalies. The data is available at https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.csv as a CSV file which you open inside Excel looks something like this:
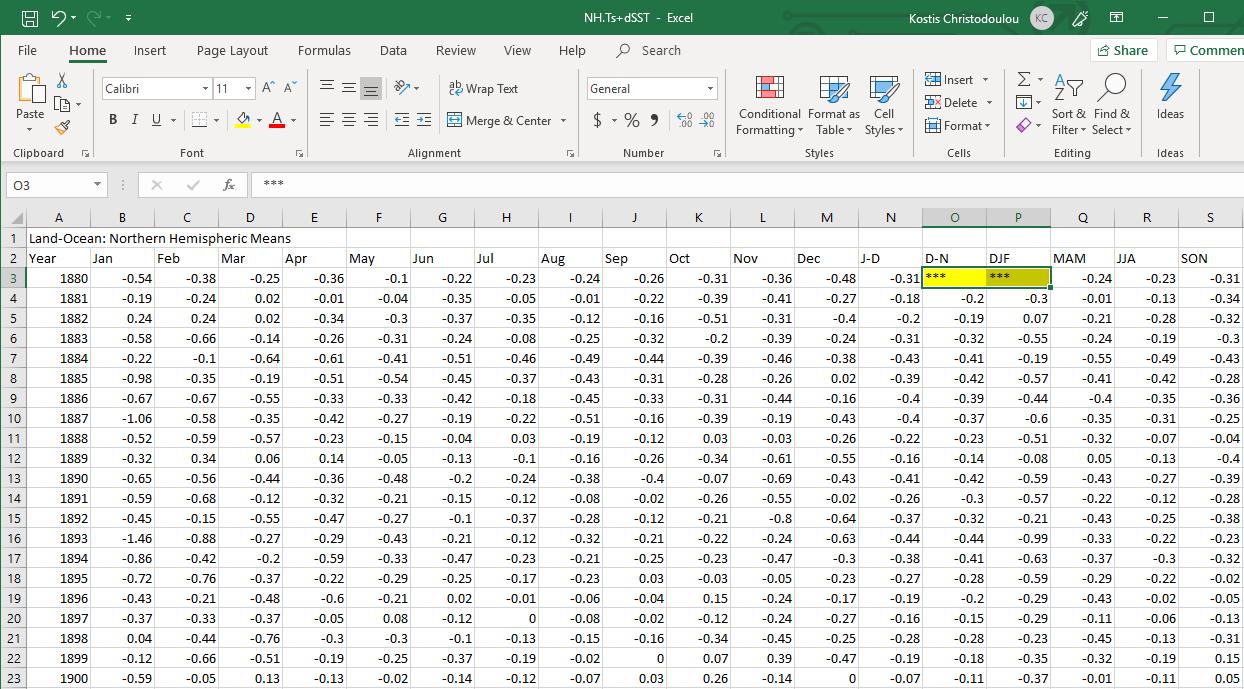
However, this is what a CSV file looks like on the inside: a bunch of values separated with commas.
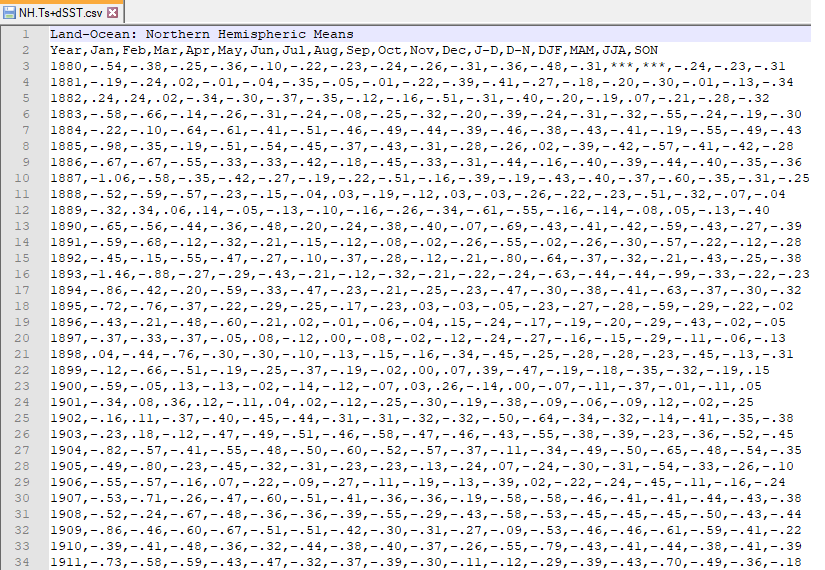
By the way, if you look at the data closely, you will notice that the
values in the D-N (December-November) and DJF
(December-January-February) columns for the year 1880 are ***. These
*** denote a missing value, in the same way that R uses the NA (or
not available) value.
If you’d like R to treat these *** values as missing, you will need to
convert them to NAs. One way to do this is to ask read_csv() to
parse *** values as NA values when it reads in the data.
Importing CSV files: read_csv()
Importing CSV is part of base R using the read.csv() command. However,
we will use the readr package and its read_csv() command that allows
us to read flat data. read_csv() is significantly (8-10 times) faster
and more user friendly than the base R command, with no need to define
rownames, no stringsAsFactors = TRUE.
Even though we only concentrate on CSV files, readr has several
functions that allow you to import a specific flat file format.
| Function | Reads |
|---|---|
read_csv() |
Comma separated values |
read_csv2() |
Semi-colon separate values |
read_delim() |
General delimited files |
read_fwf() |
Fixed width files |
read_log() |
Apache log files |
read_table() |
Space separated files |
read_tsv() |
Tab delimited values |
Just as you can import data, readr allows you to export data and save
it locally. These functions are similar to the read_ functions and
each save a tibble (or data frame) in the specific file format.
| Function | Writes |
|---|---|
write_csv() |
Comma separated values |
write_excel_csv() |
CSV that you plan to open in Excel |
write_delim() |
General delimited files |
write_file() |
A single string, written as is |
write_lines() |
A vector of strings, one string per line |
write_tsv() |
Tab delimited values |
To use a write_ function, first give it the name of the data frame to
save, then give it a filename to save in your working directory.
Importing CSV files directly off the internet
If the CSV exists on the internet and you have the URL address, you don’t have to download it to your local machine and then import it; you can import it directly off the web using the URL link.
url <- "https://data.giss.nasa.gov/gistemp/tabledata_v4/NH.Ts+dSST.csv"
weather <- read_csv(url, skip = 1, na = "***")When using the read_csv() function, we added two options:
skip = 1option is there as the real data table only starts in Row 2, so we need to skip one row.na = "***"option informs R how missing observations in the spreadsheet are coded. As discussed earlier, it is best to specify NA values here, as otherwise some of the data may not be recognized as numeric data.
Notice that the code above saves the output to an object named
weather. You must save the output of read_csv() to an object if you
wish to use it later; otherwise, read_csv() will just print the
contents of the data set at the command line.
Also, the assignment statement doesn’t produce any output for weather
because assignments don’t display anything. If we want to check that our
data has been loaded, we can see glimpse the structure of the dataframe
using glimpse() or see its contents by just typing its name:
weather.
glimpse(weather)shows us the number of observations and variables, and then, for each variable, shows the variable type; in our case, all of the variables aredblor double, namely numeric variables.weather, just invoking the name of the dataframe, shows the contents of the dataframe in the tabular form it is saved in
glimpse(weather)## Rows: 142
## Columns: 19
## $ Year <dbl> 1880, 1881, 1882, 1883, 1884, 1885, 1886, 1887, 1888, 1889, 1890~
## $ Jan <dbl> -0.36, -0.30, 0.27, -0.57, -0.17, -1.01, -0.74, -1.08, -0.48, -0~
## $ Feb <dbl> -0.51, -0.22, 0.22, -0.66, -0.09, -0.45, -0.83, -0.70, -0.60, 0.~
## $ Mar <dbl> -0.23, -0.04, 0.02, -0.15, -0.63, -0.24, -0.72, -0.44, -0.64, -0~
## $ Apr <dbl> -0.30, 0.00, -0.32, -0.29, -0.60, -0.49, -0.37, -0.38, -0.22, 0.~
## $ May <dbl> -0.06, 0.03, -0.25, -0.24, -0.36, -0.59, -0.33, -0.25, -0.16, -0~
## $ Jun <dbl> -0.16, -0.33, -0.30, -0.12, -0.43, -0.45, -0.37, -0.20, -0.03, -~
## $ Jul <dbl> -0.18, 0.08, -0.27, -0.03, -0.43, -0.34, -0.15, -0.24, 0.00, -0.~
## $ Aug <dbl> -0.26, -0.04, -0.14, -0.21, -0.49, -0.41, -0.43, -0.54, -0.21, -~
## $ Sep <dbl> -0.23, -0.27, -0.24, -0.31, -0.44, -0.40, -0.33, -0.20, -0.19, -~
## $ Oct <dbl> -0.32, -0.44, -0.52, -0.16, -0.45, -0.37, -0.31, -0.48, -0.04, -~
## $ Nov <dbl> -0.43, -0.37, -0.33, -0.42, -0.58, -0.39, -0.39, -0.27, -0.01, -~
## $ Dec <dbl> -0.40, -0.23, -0.67, -0.15, -0.47, -0.12, -0.21, -0.43, -0.24, -~
## $ `J-D` <dbl> -0.29, -0.18, -0.21, -0.27, -0.43, -0.44, -0.43, -0.44, -0.23, -~
## $ `D-N` <dbl> NA, -0.19, -0.17, -0.32, -0.40, -0.47, -0.42, -0.42, -0.25, -0.1~
## $ DJF <dbl> NA, -0.31, 0.08, -0.63, -0.13, -0.65, -0.56, -0.66, -0.51, -0.07~
## $ MAM <dbl> -0.20, 0.00, -0.18, -0.22, -0.53, -0.44, -0.47, -0.36, -0.34, 0.~
## $ JJA <dbl> -0.20, -0.09, -0.24, -0.12, -0.45, -0.40, -0.32, -0.33, -0.08, -~
## $ SON <dbl> -0.33, -0.36, -0.36, -0.30, -0.49, -0.38, -0.35, -0.32, -0.08, -~weather## # A tibble: 142 x 19
## Year Jan Feb Mar Apr May Jun Jul Aug Sep Oct Nov Dec
## <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 1880 -0.36 -0.51 -0.23 -0.3 -0.06 -0.16 -0.18 -0.26 -0.23 -0.32 -0.43 -0.4
## 2 1881 -0.3 -0.22 -0.04 0 0.03 -0.33 0.08 -0.04 -0.27 -0.44 -0.37 -0.23
## 3 1882 0.27 0.22 0.02 -0.32 -0.25 -0.3 -0.27 -0.14 -0.24 -0.52 -0.33 -0.67
## 4 1883 -0.57 -0.66 -0.15 -0.29 -0.24 -0.12 -0.03 -0.21 -0.31 -0.16 -0.42 -0.15
## 5 1884 -0.17 -0.09 -0.63 -0.6 -0.36 -0.43 -0.43 -0.49 -0.44 -0.45 -0.58 -0.47
## 6 1885 -1.01 -0.45 -0.24 -0.49 -0.59 -0.45 -0.34 -0.41 -0.4 -0.37 -0.39 -0.12
## 7 1886 -0.74 -0.83 -0.72 -0.37 -0.33 -0.37 -0.15 -0.43 -0.33 -0.31 -0.39 -0.21
## 8 1887 -1.08 -0.7 -0.44 -0.38 -0.25 -0.2 -0.24 -0.54 -0.2 -0.48 -0.27 -0.43
## 9 1888 -0.48 -0.6 -0.64 -0.22 -0.16 -0.03 0 -0.21 -0.19 -0.04 -0.01 -0.24
## 10 1889 -0.27 0.3 -0.03 0.16 -0.04 -0.07 -0.08 -0.21 -0.31 -0.42 -0.62 -0.55
## # ... with 132 more rows, and 6 more variables: J-D <dbl>, D-N <dbl>,
## # DJF <dbl>, MAM <dbl>, JJA <dbl>, SON <dbl>When you use read_csv(), read_csv() tries to match each column of
input to one of the basic data types in R. In our case, read_csv()
misidentified the contents of the Year column to a real number, rather
than an integer. You can correct this with R’s as.integer() function,
or you can read the data in again, this time instructing read_csv() to
parse the column as integers.
To do this:
- add the argument
col_typestoread_csv() - set equal the
col_typesargument to a list. - add a named element to the list for each column you would like to manually parse; in our case, we want to make column ‘Year’ an integer.
weather <- read_csv(url, skip = 1, na = "***",
col_types = list(Year = col_integer()))To complete the code, set Year equal to one of the functions below,
each function instructs read_csv() to parse Year as a specific type
of data.
| Type function | Data Type |
|---|---|
col_character() |
character |
col_date() |
Date |
col_datetime() |
POSIXct (date-time) |
col_double() |
double (numeric) |
col_factor() |
factor |
col_guess() |
let readr guess (default) |
col_integer() |
integer |
col_logical() |
logical |
col_number() |
numbers mixed with non-number characters |
col_numeric() |
double or integer |
col_skip() |
do not read this column |
col_time() |
time |
Importing CSV files saved locally
If you want to read a CSV file you have saved locally in your computer,
you must let RStudio know which folder the file lives in; in more
technical terms, you have to set the Working Directory. You can
determine the location of your working directory by running getwd().
You can change the location of your working directory by going to
Session > Set Working Directory in the RStudio IDE menu bar or use
the Ctrl+Shift+H shortcut in Windows, Cmd+Shift+H in Mac to browse
for the folder where the file resides.
Need for speed: Enter data.table::fread() and vroom::vroom()
If you have a file which is fairly large, the fread() function from
the data.table package, data.table::fread(), and vroom::vroom()
can make your life easier. You use them in a similar way to
read_csv(), but they are faster.
The following table compares how long it takes to read 22.4 million rows
from CDC’s COVID-19 Case Surveillance Public Use
Data,
with data collected as of 30 April 2021. We downloaded the CSV locally
and then read it with base R read.csv(), readr::read_csv(),
data.table::fread(), and vroom::vroom().
library(microbenchmark)
mbm = microbenchmark(
baseR = read.csv("COVID-19_Case_Surveillance_Public_Use_Data.csv"),
readr = read_csv("COVID-19_Case_Surveillance_Public_Use_Data.csv"),
data.table = fread("COVID-19_Case_Surveillance_Public_Use_Data.csv"),
vroom = vroom("COVID-19_Case_Surveillance_Public_Use_Data.csv"),
times=10
)
mbm| Unit: seconds | ||||||||
|---|---|---|---|---|---|---|---|---|
| expr | min | lq | mean | median | uq | max | neval | cld |
baseR::read.csv() |
83.24 | 85.60 | 89.71 | 89.01 | 94.26 | 96.95 | 10 | d |
readr::read_csv() |
47.09 | 48.10 | 49.69 | 49.72 | 50.11 | 54.58 | 10 | c |
data.table::fread() |
11.72 | 13.33 | 16.21 | 13.87 | 20.93 | 24.99 | 10 | b |
vroom::vroom() |
5.71 | 6.01 | 8.28 | 7.738 | 10.99 | 12.49 | 10 | a |
The difference in speed is remarkable; base R is the slowest with an
average loading time of about 90 seconds. read_csv() yields an average
of 50 seconds, data.table:fread() reduces loading time to 16 seconds,
and vroom::vroom() takes on average about 8 seconds.
Other data formats
The readr package provides efficient functions for reading and saving
common flat file data formats. For other data types, consider using :
| Package | Reads |
|---|---|
| readxl | Excel files (.xls, .xlsx) |
| haven | SPSS, Stata, and SAS files |
| jsonlite | json |
| xml2 | xml |
| httr | web API’s |
| rvest | web pages (web scraping) |
| DBI | databases |
| sparklyr | data loaded into spark |
rio: a swiss-army knife for data input-output
A really neat package to handle importing- exporting data is rio whose
authors call it A Swiss-Army knife for data input-output. It works by
determining the data structure from the file extension, uses reasonable
defaults for data import and export (e.g., ‘stringsAsFactors=FALSE’),
supports web-based import (including from SSL/HTTPS). It also has a
useful function, ‘convert()’, that provides a simple method for
converting between file types. You can read more about rio
here
Never work directly on the raw data
In 2012 Cecilia Giménez, an 83-year-old widow and amateur painter, attempted to restore a century-old fresco of Jesus crowned with thorns in her local church in Borja, Spain. The restoration didn’t go very well, but, surprisingly, the botched restoration of Jesus fresco miraculously saved the Spanish Town.
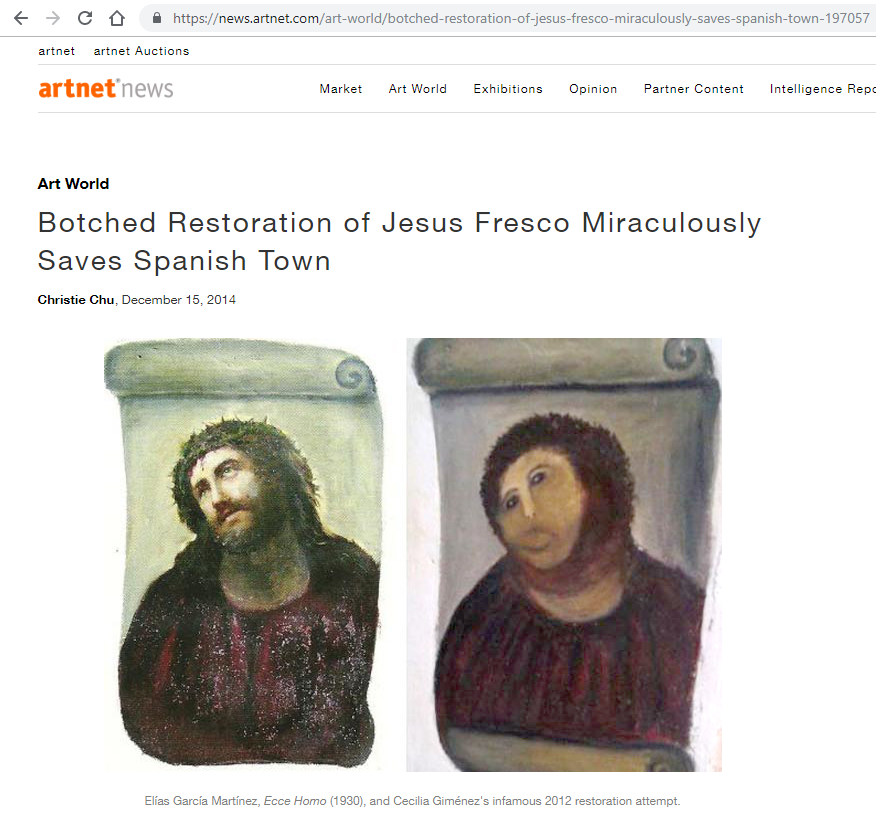
As a most important rule, please do not work on the raw data; it’s
unlikely you will have Cecilia Giménez’s good fortune to become
(in)famous for your not-so-brilliant work. Make sure you import the data
in R, leave the raw data aside, and if you make any changes tidying and
wrangling your data, save it using write_csv() with a different file
name.